Most DNA synthesis services try to screen their orders against hazardous pathogens. Separately, there are emerging techniques that can physically obfuscate genetic sequences to protect the intellectual property of synthetic genetic designs
(Purcell et al. 2018). Do these obfuscation techniques present a threat to screening systems?
I considered this question as a Summer Research Fellow in the Stanford Existential Risk Initiative. I've linked my full paper here, as well as a short summary below. The full paper requires a polish and edit, but readers might find particular sections interesting.
Bottom line up front: currently published obfuscation techniques don't appear to be feasible for obscuring hazardous genomes from screening. However, plausible future developments may disrupt this security.
Summary
Further Background
Following on from our question, the concern is that if obfuscation methods work against screening, they might enable a malicious or reckless actor to order a customer-encrypted version of a hazardous gene construct from a company, without tripping their screening systems. Then the printed construct could be 'de-obfuscated' into the genetic sequence of a dangerous pathogen.
I considered two obfuscation techniques that have been shown to work in vitro for simple synthetic designs ('circuits'): scrambling with recombinases and camouflaging with CRISPR. Scrambling is the introduction of specifically targeted inversions or excisions of subsequences, via enzymes called recombinases. Nesting and combining orthogonal recombinase sets allows a form of 'encryption' of a synthetic design - you need to know which recombinases to apply in which order to regain the original design.
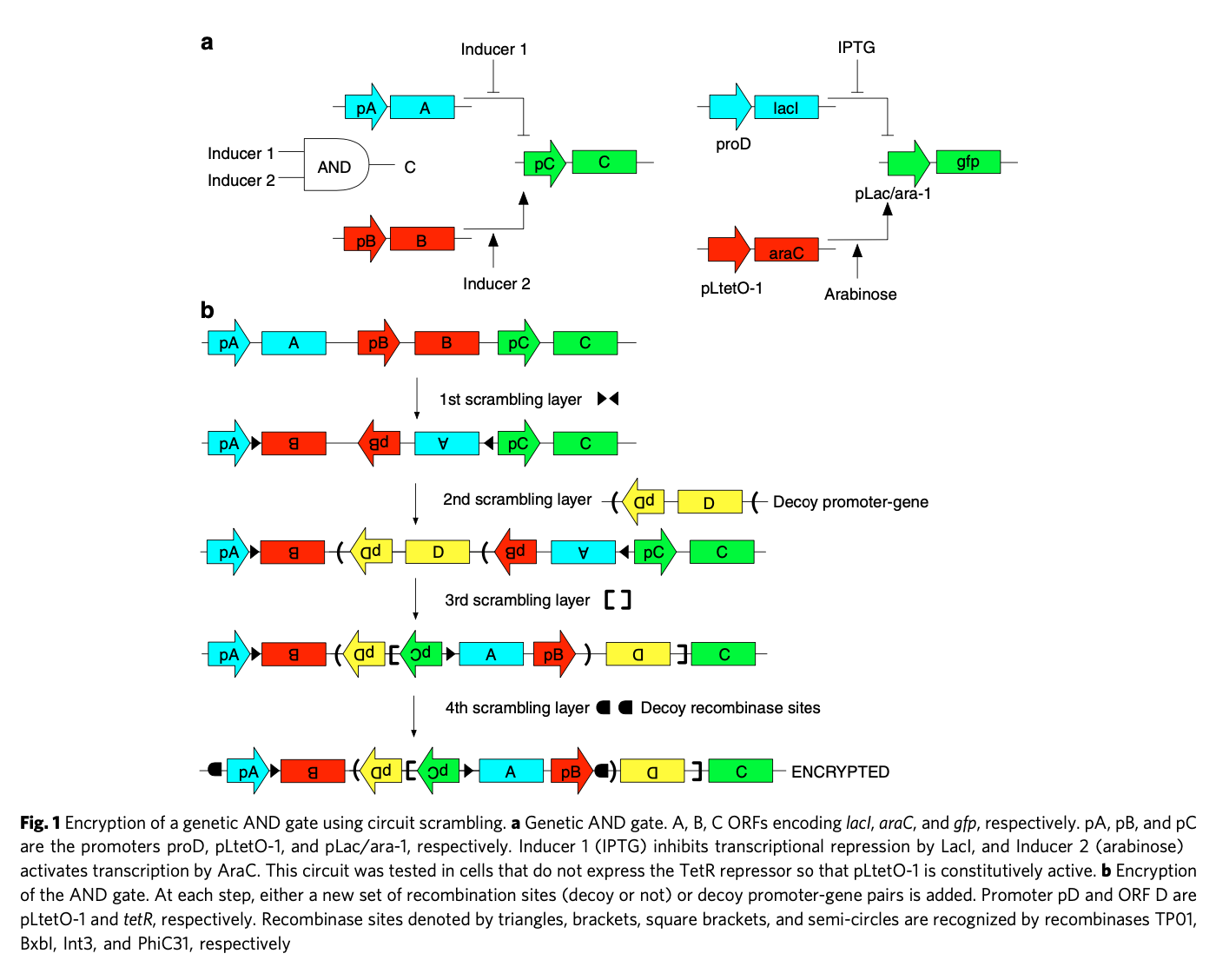
Camouflage is the introduction of 'dummy components' to obscure the true circuit design. The 'camouflaged' sequence will be difficult to interpret and not fully functional. To regain the original sequence, you must know which components are false and then subtract them with specific CRISPR keys. It is inspired by similar methods used to obscure microchip design against snooping via top-down microscopy.
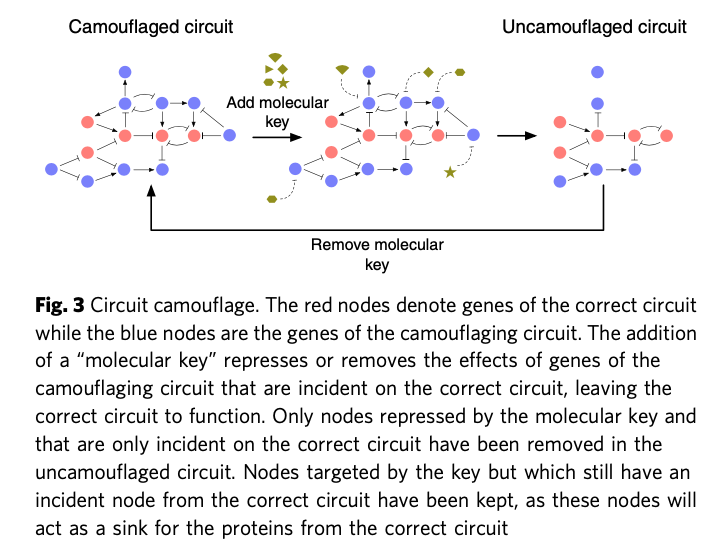
I analysed how these techniques might evade three families of screening systems: basic bioinformatics, 'random adversarial thresholds' (SecureDNA, 2021) and IARPA's 'Functional Genomic and Computational Assessment of Threats' (FunGCAT). I leave specific details of these screening approaches to the main paper.
Results
A theoretical model of the problem, combined with exploratory empirical testing, suggests that scrambling cannot feasibly obfuscate pathogen genomes against any of the three screening approaches. Even for very short genomes, the number of required recombinations greatly exceeds the set of currently discovered orthogonal recombinases. The number of required recombinations also introduces severe inefficiencies, potential interruptions to coding regions, and a significant proportion of recombination sites relative to the construct genome, while remaining vulnerable to basic investigatory follow-up.
Interestingly, if recombinase technology was to continue to advance linearly, it may present a problem specific to the 'random adversarial threshold' approach in the near-future. This is because the combinatorial possibilities of heavily scrambled sequences might outstrip the feasible size of variant databases. Further investigation might be warranted, resulting in either reassurance or adaptive response.
A similar model and argument suggests that camouflage is generally infeasible as a method to obfuscate pathogen genomes. There are key differences between circuit designs and genome identities in how their relevant information is embedded in sequences. I discuss these differences, and why they make the infeasibility of camouflage unsurprising.
A paper in Nature Biotechnology actually used camouflage to evade a basic screening process (Puzis et al. 2020). While this case does merit some concern, the hazardous sequence they ordered was extremely short - a conotoxin around 120 base pairs long. I think there is a large gulf between this case and the feasibility of camouflaging larger constructs (e.g. 1700 base pairs, the shortest RNA virus genome), though more analysis would be welcome. This problem overlaps with the problem of screening very short sequences, which could be stitched together to make a hazardous construct. Camouflage techniques may offer a different route to defeating screening, but at the cost of introducing further steps with attendant difficulties and inefficiencies. It's ambiguous if camouflage substantially alters the risk from very short sequences.
After assessing current obfuscation techniques and screening approaches, I then tried to assess how the strategic situation might change in the future. I suggested a list of possible warning signs - developments that may disrupt our current security against obfuscation.
These warning signs are:
- synthetic biology develops in such a way that valuable intellectual property is instantiated in very short sequences (10s of base pairs);
- it becomes normal for obfuscation to be commonly used;
- large (orders of magnitude) advances in CRISPR ease, capacity, and efficiency;
- obfuscation in situ (obfuscation that retains biological function in an 'encrypted' state);
- arbitrarily orthogonal recombination;
- seamless recombination; and
- diffusion of genetic recoding.
(Note: Some are these are too vague for my liking. Creating a better list and sharpening its items might be a useful project.)
Information hazards
Finally, this case is an interesting example of managing information hazards 'beyond the openness-secrecy axis' (Lewis et al. 2018). I was initially quite worried about the potential information hazards of the project, as I was unsure of the likelihood of discovering live vulnerabilities to obfuscation in current screening systems. Early conversations with my advisor were reassuring that current vulnerabilities were extremely unlikely, though I still proceeded with caution. I avoided analysing techniques and protocols that weren't widely published or accessible, and I generally aimed to frame my research process to focus on 'what would a screening protocol need to do?' rather than 'what is a viable evasion strategy?'
The potential vulnerabilities I did discuss were conjectured for either screening protocols that are still in the development stage (where red-teaming can be valuable) or for obfuscation techniques that aren’t currently feasible (where foresight is useful) (Zhang & Gronvall 2018).
Acknowledgements
Thanks to the Stanford Existential Risk Initiative's tireless staff and passionate co-fellows. Thanks to my mentor Dr. Michael Montague. Thanks to the Future of Humanity Institute, which I visited as an intern in early 2020 and worked on a related, unfinished project. That project gave me the background to propose and tackle this project. Thanks to Dr. Cassidy Nelson who mentored me on that internship, and Dr. Gregory Lewis who initially suggested steganography as a potentially interesting sub-topic in the screening problem.
References (incomplete, full list in paper)
Lewis, Gregory, Piers Millett, Anders Sandberg, Andrew Snyder‐Beattie, and Gigi Gronvall. 2018. "Information Hazards In Biotechnology". Risk Analysis 39 (5): 975-981. doi:10.1111/risa.13235.
Purcell, Oliver, Jerry Wang, Piro Siuti, and Timothy K. Lu. 2018. "Encryption And Steganography Of Synthetic Gene Circuits". Nature Communications 9 (1). doi:10.1038/s41467-018-07144-7.
Puzis, Rami, Dor Farbiash, Oleg Brodt, Yuval Elovici, and Dov Greenbaum. 2020. "Increased Cyber-Biosecurity For DNA Synthesis". Nature Biotechnology 38 (12): 1379-1381. doi:10.1038/s41587-020-00761-y.
SecureDNA. 2021. “Random adversarial threshold search enables specific, secure, and automated DNA synthesis screening” (preprint). securedna.org Available from: https://www.securedna.org/download/Random_Adversarial_Threshold_Screening.pdf (Accessed 1/7/21)
Zhang, Lisa, and Gigi Kwik Gronvall. 2018. "Red Teaming The Biological Sciences For Deliberate Threats". Terrorism And Political Violence 32 (6): 1225-1244. doi:10.1080/09546553.2018.1457527.
