Supervisor: Akhil Bansal
Executive Summary
Biosurveillance systems are needed to mitigate against three primary threats. First, the accidental release of pathogens from labs. Secondly, the deliberate release of pathogens from bad actors with malicious intent. Lastly, the natural occurrence of pathogens that have the potential to create epidemics and pandemics. A robust biosurveillance system would aim to identify and monitor new and known pathogens with the goal of reducing the severity of the spread of the infectious diseases. From the COVID-19 pandemic, it’s clear that modern biosurveillance systems have the potential to improve tremendously. To do so, a better understanding of the bottlenecks biosurveillance systems face is needed.
The report consists of two standalone parts. This is part I of the report that introduces the current pathogen biosurveillance landscape, with a focus on biosurveillance technologies. Part II of the report explores the potential and bottlenecks of three technologies commonly used for biosurveillance - Polymerase Chain Reaction (PCR), Loop-Mediated Isothermal Amplification (LAMP) and Metagenomics Sequencing.
Part I of this report will be most useful for you if you would like to be informed about the overall biosurveillance landscape, the different types of biosurveillance systems, and the main types of technologies used in these biosurveillance landscapes. If you would like to have a deeper understanding of PCR, LAMP, and metagenomics sequencing and how they can be applied in biosurveillance, please view Part 2 of the report.
Throughout the project, we used a combination of empirical research and expert interviews to gather information. A list of the organizations that we have talked to is included in Appendix 2. In total, we gathered the opinions of 25 experts and sourced over fifty peer-reviewed research papers.
Chapter 1 - Prioritizing Biosurveillance Systems (link)
In the first chapter of the project, we developed a weighted factor model (details of the WFM) to assess the robustness of the existing biosurveillance systems. The aim of the WFM is to prioritize a few biosurveillance systems that we believe are the most important. The WFM assessed ten aspects of a biosurveillance system, including its usefulness, feasibility and potential risks. For every surveillance system, participants could adjust the weighting of each criteria and assign a score to each criteria for the relevant surveillance system. The final WFM incorporated the output of seven experts. A higher score in the WFM model indicated a higher relevance of that biosurveillance system to our project.
Types of Surveillance Systems | PoP | Clinical | Environmental | Animal | Syndromic | Digital |
| Averaged score | 4.8 | 4.9 | 5.6 | 4.6 | 4.3 | 4.9* |
Table 1 - Summary of results from the Weighted Factor Model. We identified Point-of-Person (PoP), Clinical and Environmental Surveillance systems as the most relevant for our project.
* Despite Digital Surveillance having a high score, we decided not to focus on it for the reason mentioned in insight 3 below.
Some further key insights gathered from the WFM are shown below and linked to further explanations.
Insight 1– Point-of-Person and Clinical Surveillance can benefit largely from technological developments
Insight 2 – Environmental surveillance is a very promising area
Insight 3– The main challenges Syndromic and Digital Surveillance face are data- and operation-related
Insight 4 – Zoonotic Surveillance cannot easily be used for early-stage human pathogen detection
We identified a list of technologies commonly used in PoP, Clinical, and Environmental Surveillance across the sampling, concentration, extraction, detection and quantification stages. Due to the ten-week time constraint of the project, we narrowed our focus to Polymerase Chain Reaction (PCR), Loop-Mediated Isothermal Amplification (LAMP) and Metagenomic Sequencing. For each of these technologies, we reviewed their current states, limitations, and potential avenues for improvements.
Chapter II - Overview of Technologies used in Biosurveillance
In the second chapter, we categorized the main technologies used at different stages of biosurveillance, from sampling, concentration, extraction, detection, to quantification. The list of technologies were made after two weeks of empirical research and talking to experts. We also provided a framework to assess the importance of each of these technologies. However, we were unable to apply this criteria rigorously to assess the technologies we identified, primarily due to lack of information that we can gather. It was difficult to find experts to interview for each of these technologies in the time limit of our research fellowship, and we couldn’t find existing research papers that ranked the importance of these technologies in the way we wanted.
In the end, we used a qualitative and intuitive assessment of how important these technologies were and decided on PCR, LAMP, and metagenomics sequencing. It would be useful to conduct a more rigorous review of these technologies if resources are available, as different conclusions may be reached.
Acknowledgment
We would like to thank CERI (Cambridge Existential Risk Initiatives) and the biosecurity team for helping us make this happen during the 2022 Summer Research Fellowship. In particular, we are extremely grateful for our supervisor Akhil Bansal and biosecurity lead Dewi Erwan.
Table of Contents
Chapter I - Introduction to Biosurveillance Systems
- The Need for a Biosurveillance System
- Different Types of Biosurveillance Systems
- Prioritizing Surveillance Systems
- Insight 1 – Point-of-Person and Clinical Surveillance can benefit largely from technological developments
- Insight 2 – Environmental surveillance is a very promising area
- Insight 3 – The main challenges Syndromic and Digital Surveillance face are data- and operation-related
- Insight 4 – Zoonotic Surveillance cannot easily be used for early-stage human pathogen detection
- Bottlenecks of Surveillance Systems
Chapter II - Biosurveillance Technologies
Chapter I - Introduction to Biosurveillance Systems
The Need for a Biosurveillance System
In the twenty-first century, it’s become increasingly apparent that humanity is in dire need of a robust biosurveillance system. A biosurveillance system is characterized by identifying novel and known pathogens in an environment, this includes living organisms such as plants and humans, water systems and more. Biosurveillance systems serve to monitor the spread of diseases across these populations and environments so they can be contained. There has been an increase in zoonotic infectious diseases in this century. Notably, the 2003 SARS Outbreak (WHO, 2015), the 2009 Swine Flu Pandemic (CDC, 2012), the 2012 Middle East Respiratory Syndrome Coronavirus outbreak (ECDC, 2021), the 2013-2016 Ebola epidemic (CDC, 2014), the 2015 Zika Virus epidemic (WHO, 2015) and recently the Covid-19 pandemic. As of the time of writing this report, the Monkeypox outbreak is increasingly worrying. In the future, the frequency and source of pandemics are likely to increase.
As labs work to better understand the underlying mechanisms of pathogen mutations, the possibility of accidental releases of highly transmissible and deadly pathogens through water drainage, air vents and other means become increasingly worrying. Furthermore, as more DNA synthesis companies enable the purchasing of synthetic DNA, it’s become easier for bad actors to engineer pathogens that are highly transmissible and fatal. Engineered pandemics pose a serious threat to humanity and have the potential to be Global Catastrophic Biological Risks (GCBRs). The Center for Health Security defines a GCBR as a threat that “could lead to sudden, extraordinary, widespread disaster beyond the collective capability of national and international governments and the private sector to control. If unchecked, GCBRs would lead to great suffering, loss of life, and sustained damage to national governments, international relationships, economies, societal stability, or global security” (Alexopulos, 2019).
A biosurveillance system is particularly useful in the early detection of new and known infectious diseases, predicting the spread of diseases and helping contain pathogens before they impair the lives of potentially millions of people. The ideal biosurveillance system would involve rapid early detection of novel and known pathogens allowing relevant actors to identify, contain and mitigate any biological risk. The implementation of a robust, integrated biosurveillance system in this decade will better prepare humanity for the emerging threats of the twenty-first century and onwards.
The recent Covid-19 pandemic furthers the imperative to develop robust biosurveillance systems to prepare humanity for future, more severe pandemics. There is increased interest in preventing pandemics due to the Covid-19 pandemic but this interest is diminishing. Therefore, immediate action must be taken by relevant actors soon. The next pandemic should not be humanity’s wake-up call to better prepare for the future.
Different Types of Biosurveillance Systems
To improve upon current biosurveillance efforts, we must first understand the types of biosurveillance systems and their functions.
| Type of Surveillance | Function | Currently Used Example |
| Point of Person | Diagnostics that do infectious disease detection at the patient level without further equipment. | Covid-19 Rapid Lateral Flow Test Uses a sample of bodily fluids to detect viral proteins without the use of complex laboratory equipment. |
| Clinical | Diagnostics that do infectious disease detection at the point of care. This could involve taking a sample at the clinic, sending it back to the lab and receiving a result a week later. | A lab technique that enables the detection of viruses in a sample. |
| Sentinel | Monitoring infectious diseases among high vulnerability people such as health care workers, TSA agents, and livestock workers. | Belgium’s Sentinel System. Aims to do early detection of Work Related Disease by allowing physicians to report diseases caused by a patient’s occupation. |
| Environmental | Monitoring and detection of known and unknown pathogens in wastewater systems, and busy public spaces like airports or farms. | CDC COVID Wastewater Surveillance Wastewater from sewershed collected and sent to laboratories for SARS-CoV02 testing |
| Human / Animal Monitoring | Monitoring and detection of pathogens in livestock, wild animals and humans through wearable devices and frequent sampling. | An integrated approach to monitor health of humans, domestic and wild animals, and the wider ecosystems. |
| Syndromic | Monitoring of symptoms of infectious diseases before a diagnosis is made. | Tracks symptomatic cases in the UK |
| Digital | Tools that involve people reporting cases to a larger database and extracting insights from these databases. | Users infected with Covid-19 report their location and app notifies people who were close by (Canada) |
Table 2 - Summary of the different biosurveillance systems and examples of used cases.
Prioritizing Surveillance Systems
We developed a Weighted Factor Model (details of our model) to assess and compare the robustness of the different surveillance systems described above. Our WFM included ten factors that include the feasibility and usefulness of the surveillance systems, as well as the quality of evidence that exists. This model is chosen primarily for its ability to incorporate many factors and produce a quantitative number for each of the surveillance systems for systematic comparisons. One potential drawback to the model is that some of the factors are rather subjective. To obtain more comprehensive results, we gathered responses from 7 experts in the field. The updated view is presented below.
The specific criteria used in the WFM are included in Appendix 1. Scores ranging from 1 to 7 are assigned for each factor for every surveillance system, and the total score is calculated based on the weightings assigned to each criterion. Point-of-Person, Clinical, and Environmental Surveillance scored the highest priority according to our WFM, therefore we will focus on these areas in the remainder of this report.
Insight 1 – Point-of-Person and Clinical Surveillance can benefit largely from technological developments
Clinical Surveillance is one of the most obvious ways of pathogen detection and the system is largely in place. However, the quality and time needed between symptom emergence and diagnosis depend highly on the equipment and personnel available at the clinics. Polymerase Chain Reaction (PCR) is the golden standard for pathogen detection due to its high sensitivity and specificity, but the equipment is costly (thousands to tens of thousands of USD) and the operation process is complicated, requiring skilled personnel to perform the tests. There is therefore a demand for alternative tests that are cheaper and easier to operate. Furthermore, PCR test is a form of targeted diagnosis and is unable to detect novel pathogens. There is increasing attention to develop pathogen-agnostic tests that can detect both existing and emerging pathogens.
Other drawbacks of clinical surveillance is its difficulty in detecting asymptomatic infections as patients would only visit hospitals when they feel sick. The act of traveling to and from the clinics also increases the risks of transmission. Point-of-Person (PoP) surveillance can overcome these problems by having people regularly take diagnostic tests at home. PoP devices in the form of LFTs proved their effectiveness during the COVID-19 pandemic. Ideally, we would like to have similar tests that are less invasive (e.g. requiring only saliva samples), have multiplex capability so that they can detect a range of pathogens simultaneously, are pathogen agnostic with the use of metagenomic sequencing, and provide (near) real-time, easily interpretable results. These devices can be connected to large databases for integration.
Getting regulatory approval is a large bottleneck in advancing PoP surveillance (Mardis 2017), but there is now an increasing push to address this issue. For example, a joint statement between the UK government and various research organizations has called for a simplified regulatory environment (Department of Health & Social Care, 2022). A simplified regulatory framework would encourage more technological developments, therefore the current biosurveillance landscape is in favor of more technological developments in these two important surveillance systems.
Insight 2 – Environmental surveillance is a very promising area
Environmental surveillance, especially wastewater surveillance, has attracted much attention from the scientific community and government departments due to the various success stories seen in detecting pathogens. Wastewater surveillance was first implemented for tracking poliovirus in the 1990s (Asghar et al., 2014), which proved to be four to five times more sensitive in detecting outbreaks compared to syndromic surveillance. The finding that SARS-CoV RNA can be detected in human feces showed that wastewater surveillance can potentially detect a large range of pathogens. Indeed, scientists were able to capture the rise and fall of novel coronavirus cases using wastewater testing (Larsen & Wigginton, 2020). This data also aided in discovering novel strains of the virus (Venugopal et al., 2020).
One of the biggest advantages of environmental surveillance is its high efficiency, as taking a single wastewater sample could cover the population of an entire region. Some of the technological barriers to the wider adoption and broader usefulness of environmental surveillance are the need for highly sensitive testing equipment and concentration techniques that work for a variety of pathogens including bacteria, virus, and fungi. We are still at the very early stage of wastewater sampling, but there are several concrete technological bottlenecks that we can work on to better this surveillance system which will be discussed below.
Insight 3 – The main challenges Syndromic and Digital Surveillance face are data- and operation-related
The computational infrastructure for data filtering, storing, and sharing is mostly in place, and algorithms have been developed for anomaly detection. The main challenges syndromic and digital surveillance face is the lack of a centralized database between key parties and operational protocols, such as standardized ways of writing and storing data. This leads to problems such as a large variability in the amount of data available by regions, disparate data quality and a lack of interoperability between data sources. Furthermore, there is a lack of incentive alignment on the importance of data sharing between regions and nations. Although important, addressing such challenges are out of the scope of this project. The scope of this project will be discussed in more detail in Chapter II.
Insight 4 – Zoonotic Surveillance cannot easily be used for early-stage human pathogen detection
With increased globalization and animal farming, zoonotic diseases are becoming more common. Some scientists estimated that 60% of known infectious diseases are zoonotic, and it can account for up to 75% of new emerging pathogens. These zoonotic pathogens typically infect animals first, such as wildlife animals or livestock, and are transmitted to humans either directly via mediums or indirectly via vectors.
However, while we can use animal surveillance systems to regularly monitor animals’ health and collect samples, we lack the capability of characterizing the pathogens effectively so as to predict their virulence and transmissibility to humans. This is a serious problem as it is infeasible to analyze and monitor every single pathogen strain found in animals. Microbiologists are working on this problem, but until then, zoonotic surveillance may not be suitable as an early-warning system.
Bottlenecks of Surveillance Systems
Point of Person Surveillance
The ideal PoP diagnostics would be pathogen agnostic and provide rapid, on-site results of all the pathogens in a patient’s system. These tests would be taken frequently and by the majority of the population so data could be streamlined and used for monitoring outbreaks.
In comparison, PoP diagnostics today primarily include rapid antigen tests and rapid antibody tests with some CRISPR diagnostic methods like SHERLOCK (Mustafa, 2021) that test for specific pathogens like Covid-19 and HIV. Other emerging techniques include Lab on Chip (LoC) (Wu, 2018) concepts that try to automate and miniaturize benchtop PCR machines and improved SHERLOCK techniques like miSherlock (De Puig, 2021).
Today, PoP diagnostics face numerous challenges. First, the most used PoP diagnostic tools are pathogen-specific. There exists no ubiquitous pathogen-agnostic PoP diagnostic tool. Metagenomic sequencing appears to be the most promising solution to this problem but it suffers from numerous issues including its high costs, library prep methods that require skilled personnel, and long read times. These challenges are further discussed below.
Clinical Surveillance
Similar to PoP methods, an ideal clinical surveillance system would involve testing patients for multiple pathogens at the point of care and receiving a rapid, on-site result.
In clinical settings, multiplex assays are used to test for the presence of multiple pathogens but this often introduces a week of delay from sample to result time. Metagenomic sequencers are not sufficiently advanced to be used in clinical use for multiple reasons. First, they require sample preparation and library preparation in the lab by skilled professionals. Second, skilled professionals are needed to make sense of the genomic data and operate relevant equipment. Lastly, they have long read-out times and rely on incredibly expensive equipment.
In clinical settings, the primary tests used are pathogen-specific tests. Most commonly, saliva or nasal swabs are taken and sent back to labs for PCR to be run. Other commonly used molecular diagnostic tools are LAMP tests. Antibody tests include ELISA (Medline), with the most common CRISPR tests being CARMEN (Broad Institute, 2022) and SHINV2 (Arizti-Sanz, 2021).
Environmental Surveillance
An ideal environmental surveillance system could rely on frequent and automatic sampling at key locations, and combined with different sequencing methods could detect both existing and novel pathogens. The gold standard pathogen detection method is PCR (National Human Genome Research Institute, n.d.), but novel methods such as LAMP (New England Biolabs Inc, n.d.) could become more popular. Environmental surveillance provides unique advantages compared to PoP and clinical surveillance as each sample contains the sequences of many individuals. As of 2021, 55 out of 195 countries (of which 36 are high-income, 11 are upper middle-income, and 8 are lower-middle income), contain wastewater monitoring (World Health Organization, 2022).
However, in reality, this benefit is unlikely to be fully captured due to several challenges. We will focus on wastewater sampling as it is the most widely used environmental surveillance method, but there are other environmental surveillance methods that could be useful as well (Ramuta et al., 2022). First, wastewater is highly unpredictable and varies by region. Consequently, the pathogen concentration varies largely depending on the time and place of collection. Secondly, pathogens are greatly diluted in the environmental samples. This means that there may be low pathogen concentration in samples for diagnostics to run accurately. Furthermore, our understanding of microbiology is not enough to systematically estimate what proportion of pathogens would be detectable in wastewater. So far, we have used wastewater to detect specific pathogen species such as poliovirus and SARS COVID-19, but a broader understanding would be crucial before implementing globally distributed and coordinated wastewater surveillance systems.
Currently, we are only able to detect specific pathogens that are relatively concentrated in the samples, with a high probability of false negatives. The extent to which we can backtrace the diagnostic results to the pathogen concentration in the population also varies significantly depending on the labs, equipment, and methods used.
Chapter II - Biosurveillance Technologies
Narrowing our focus to PoP, clinical and environmental surveillance, we then mapped out the primary technologies needed for each of these surveillance systems, and the technological bottlenecks for each of these technologies.
Why Focus on Technological Bottlenecks
In our research, we’ve come to understand that creating effective biosurveillance systems encompasses multiple types of challenges. Below, we outlined the primary technology development stages (Diagram 1).
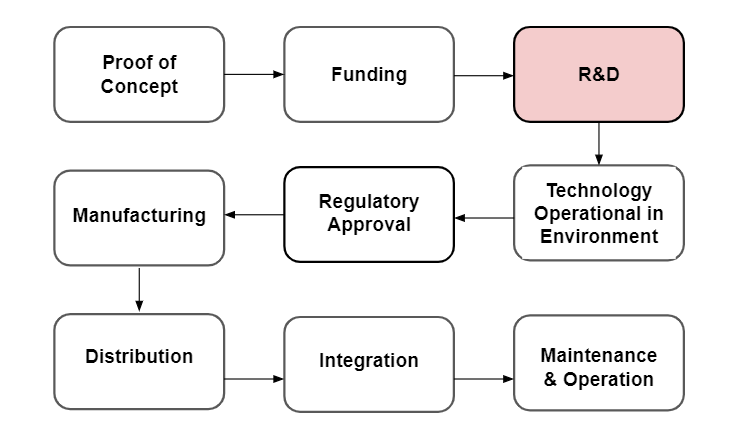
Diagram 1 - The primary technological development stages with a focus on R&D and Implementation.
While we know some of the barriers listed can significantly reduce how promising certain technologies are over others, we decided to narrow the scope of this project to technological bottlenecks that address R&D challenges. Research and development of technologies are the most fundamental steps in developing biosurveillance systems as all later challenges such as implementation require the technology to be fully functioning first. For example, our biosurveillance systems would be incredibly vulnerable to attacks from novel and known pathogens without functioning diagnostics or DNA sequencers. Fortunately, there has been an increase in pandemic prevention efforts due to the Covid-19 pandemic making it a crucial time to understand and think about how to most effectively distribute funding across focus areas.
Identifying Technologies
Through reading papers and talking to experts at the Broad Institute, the Nucleic Acid Observatory, the Future of Humanity Institute, and others, we identified a list of technologies that are used for clinical, PoP, and environmental surveillance. There might be some relevant technologies that we haven’t included here, but we are fairly confident that the list includes the most important ones. To present the information more clearly, we divided the process of pathogen surveillance (Diagram 2) into five stages: sampling, concentration, extraction of target pathogens, detection, and quantification. Point-of-Person, clinical and environmental surveillance differ largely in sampling methods, but the subsequent steps are quite similar.
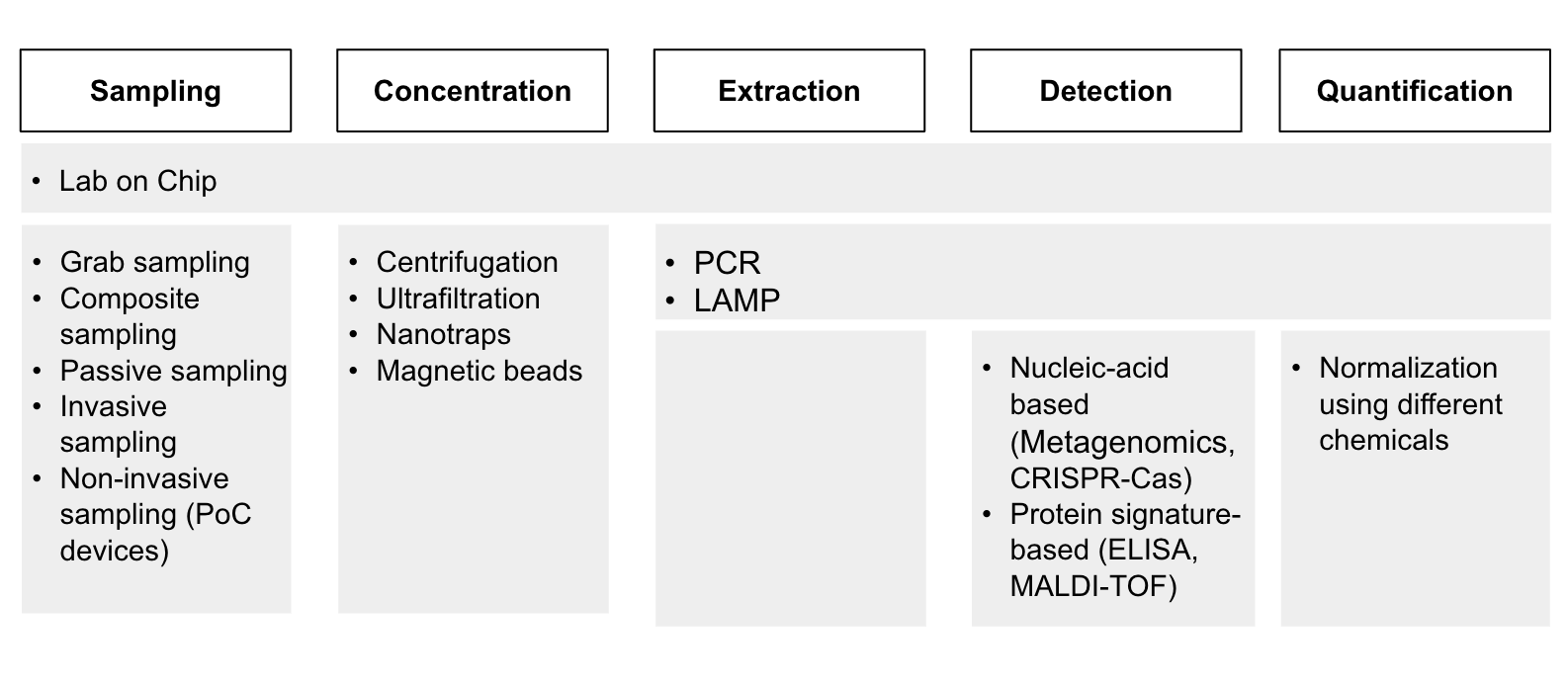
Diagram 2 - An overview of the technologies involved in PoP, clinical and environmental surveillance across the pathogen surveillance process.
Prioritizing Technologies
Below, we’ve listed the criteria we used when deciding to focus on specific technologies over others. Prioritization of technologies was primarily a qualitative ranking as many of the criteria used were difficult to quantify.
Criteria for Prioritization
In researching biosurveillance technologies, we aim to pick technologies that have the highest potential to make a large positive impact. The technologies that would naturally have the ability to do this are those that are highly accessible, inexpensive, quick, and can be used for multiple pathogens.
| Criteria For Prioritization | Description | Justification |
| Potential to be pathogen agnostic | Diagnostics can test for multiple pathogens. |
|
| Ability to be used ubiquitously | Technology can be scaled for widespread use. It is economical, manufacturing processes are sufficiently efficient. |
|
| Time to result | Time from sample taken to result. |
|
| Overlap in core technology / mechanism | Several other technologies rely on the same mechanism so if we improve the core mechanism, this benefits several other techniques. |
|
| Cost-effectiveness of the intervention at scale | **We were unable to find data or forecasts, though this will become increasingly more important (and easier to evaluate) later on in the R&D and commercialization pipeline |
|
| Potential for dual-use by bad actors | Chance that the technology could be misused by bad actors. For example, central databases that contain patient information could be hacked. |
|
| Neglectedness | Not worked on by many organizations and people |
|
| Technology Readiness Level | Ideally aiming for TRL 3-6. These are technologies that have a proof of concept and are being further validated and developed in their working environments. |
|
Table 3 - Criteria for prioritizing technologies.
Prioritized technologies
Although our intention was to apply the criteria above to assess each of the technologies, we were unable to do so primarily due to the lack of information that we can gather. It was difficult to find experts to interview for each of these technologies in time, and we couldn’t find existing research papers that ranked the importance of these technologies in the way we wanted. In the end, we used a qualitative and intuitive assessment of how important these technologies were and decided on PCR, LAMP, and metagenomics sequencing. (Diagram 3).
While we decided to put our focus on these technologies due to time constraints, we still believe that further research must be done to better understand the technological bottlenecks in the other mentioned technologies.
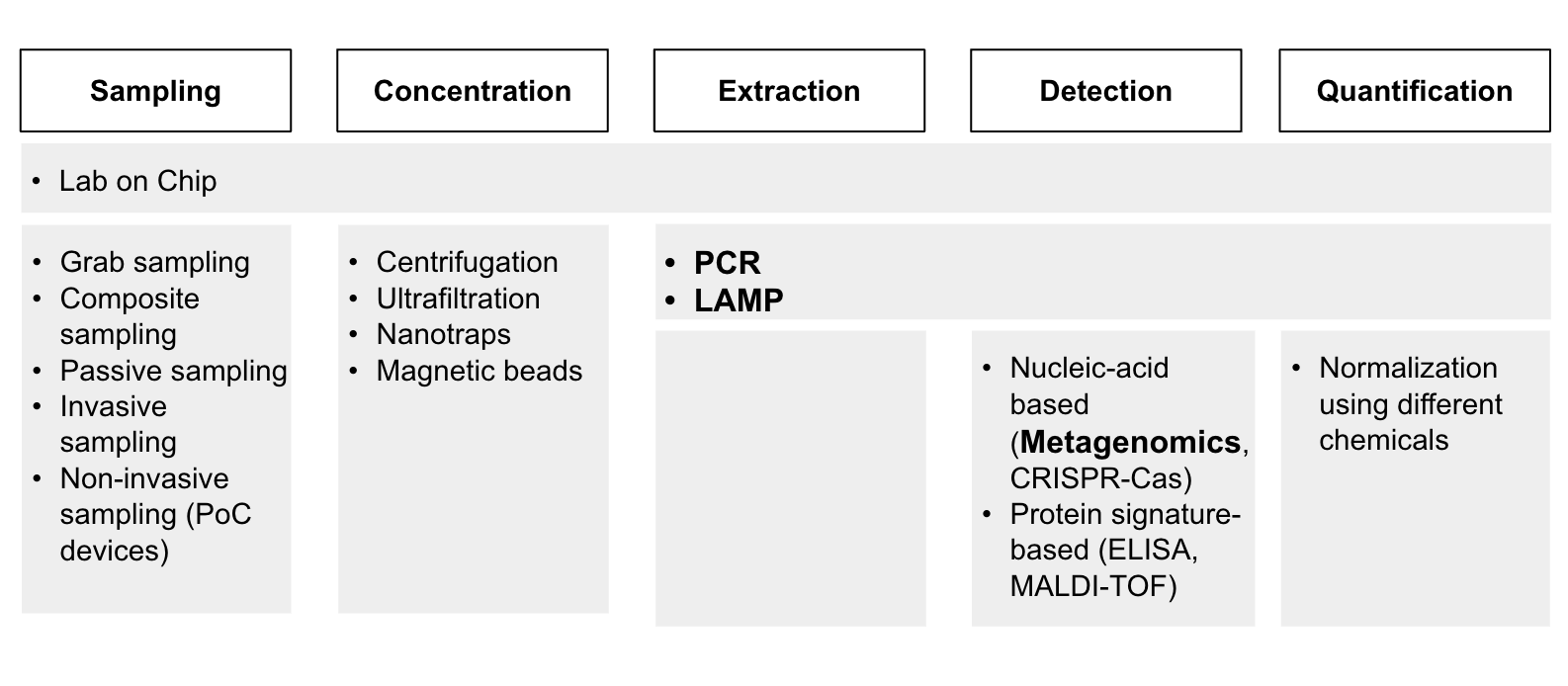
Diagram 3 - Prioritized technologies (bolded) for PoP, clinical and environmental surveillance.
References
“Alert and Sentinel Systems: Signaal, Netherlands/Belgium.” OSHA, National Human Genome Institutean biosurveillance https://osha.europa.eu/en/publications/alert-and-sentinel-systems-signaal-netherlandsbelgium.
Alexopulos, Nick. “First Working Definition of Global Catastrophic Biological Risks.” Johns Hopkins Center for Health Security, 9 Aug. 2019, www.centerforhealthsecurity.org/news/center-news/2017/2017-07-27_global-catastrophic-biological-risk-definition.html
Aridgides, L.J., et al. “Multiplex PCR Allows Simultaneous Detection of Pathogens in Ships' Ballast Water.” Marine Pollution Bulletin, vol. 48, no. 11-12, 2004, pp. 1096–1101., https://doi.org/10.1016/j.marpolbul.2003.12.017.
Arizti-Sanz, Jon, et al. “Equipment-Free Detection of SARS-COV-2 and Variants of Concern Using CAS13.” 2021, doi:10.1101/2021.11.01.21265764
Asghar, H., et al. “Environmental Surveillance for Polioviruses in the Global Polio Eradication Initiative.” Journal of Infectious Diseases, vol. 210, no. suppl 1, 2014, https://doi.org/10.1093/infdis/jiu384.
Bao, Yijuan, et al. “Cut-Lamp: Contamination-Free Loop-Mediated Isothermal Amplification Based on the CRISPR/cas9 Cleavage.” ACS Sensors, vol. 5, no. 4, 2020, pp. 1082–1091., https://doi.org/10.1021/acssensors.0c00034.
Biolabs, New England. “Loop-Mediated Isothermal Amplification.” NEB, https://international.neb.com/applications/dna-amplification-pcr-and-qpcr/isothermal-amplification/loop-mediated-isothermal-amplification-lamp.
Cai, Sheng, et al. “Phosphorothioated Primers Lead to Loop-Mediated Isothermal Amplification at Low Temperatures.” Analytical Chemistry, vol. 90, no. 14, 2018, pp. 8290–8294., https://doi.org/10.1021/acs.analchem.8b02062.
Chiu, Charles Y., and Steven A. Miller. “Clinical Metagenomics.” Nature Reviews Genetics, vol. 20, no. 6, 2019, pp. 341–355., doi:10.1038/s41576-019-0113-7.
“CRISPR-Based Diagnostic Chips Perform Thousands of Tests Simultaneously to Detect Viruses.” Broad Institute, 8 Mar. 2022, www.broadinstitute.org/news/crispr-based-diagnostic-chips-perform-thousands-tests-simultaneously-detect-viruses.
“Data Science for High Throughput Sequencing: Stanford .” Data Science for High-Throughput Sequencing, Stanford: EE 372, June 2016, data-science-sequencing.github.io/Spr2016/Spr2016/.
De Puig, Helena, et al. “Minimally Instrumented Sherlock (Misherlock) for CRISPR-Based Point-of-Care Diagnosis of SARS-COV-2 and Emerging Variants.” Science Advances, vol. 7, no. 32, 2021, doi:10.1126/sciadv.abh2944
Delahaye, Clara, and Jacques Nicolas. “Sequencing DNA with Nanopores: Troubles and Biases.” PLOS ONE, vol. 16, no. 10, 2021, doi:10.1371/journal.pone.0257521
“DNA and RNA Sequencing Kits.” Oxford Nanopore Technologies, 26 Aug. 2022, nanoporetech.com/products/kits.
“DNA Sequencing.” Oxford Nanopore Technologies, 10 June 2020, nanoporetech.com/applications/dna-nanopore-sequencing.
“ EE 372: Data Science for High-Throughput Sequencing Stanford University .” Lecture 2: Basics of DNA & Sequencing by Synthesis, data-science-sequencing.github.io/Win2018/lectures/lecture2/.
“Elisa Blood Test: Medlineplus Medical Encyclopedia.” MedlinePlus, U.S. National Library of Medicine, medlineplus.gov/ency/article/003332.html.
El-Tholotha, Mohamed, et al. “A Single and Two-Stage, Closed-Tube, Molecular Test for the 2019 Novel Coronavirus (COVID-19) at Home, Clinic, and Points of Entry.” 2020, https://doi.org/10.26434/chemrxiv.11860137.
Environmental Surveillance for SARS-COV-2 to Complement Public Health Surveillance, 2022.
Esbin, Meagan N., et al. “Overcoming the Bottleneck to Widespread Testing: A Rapid Review of Nucleic Acid Testing Approaches for COVID-19 Detection.” RNA, vol. 26, no. 7, 2020, pp. 771–783., https://doi.org/10.1261/rna.076232.120.
“First Global Estimates of 2009 H1N1 Pandemic Mortality Released by CDC-Led Collaboration.” Centers for Disease Control and Prevention, Centers for Disease Control and Prevention, 25 June 2012, www.cdc.gov/flu/spotlights/pandemic-global-estimates.html.
“Geographical Distribution of Confirmed MERS-COV Cases by Country of Infection and Year.” European Centre for Disease Prevention and Control, 7 May 2021, www.ecdc.europa.eu/en/publications-data/geographical-distribution-confirmed-mers-cov-cases-country-infection-and-year.
“Gridion.” Oxford Nanopore Technologies, 1 Mar. 2022, nanoporetech.com/products/gridion.
Head, Steven R., et al. “Library Construction for next-Generation Sequencing: Overviews and Challenges.” BioTechniques, vol. 56, no. 2, 2014, pp. 61–77., doi:10.2144/000114133.
Hsieh, Kuangwen, et al. “Simultaneous Elimination of Carryover Contamination and Detection of DNA with Uracil-DNA-Glycosylase-Supplemented Loop-Mediated Isothermal Amplification (UDG-Lamp).” Chemical Communications, vol. 50, no. 28, 2014, p. 3747., https://doi.org/10.1039/c4cc00540f.
“Illumina DNA Prep.” Illumina DNA Prep | Flexibility for Many Whole-Genome Sequencing Applications, www.illumina.com/products/by-type/sequencing-kits/library-prep-kits/nextera-dna-flex.html
“Joint Statement from the UK Government, CEPI, IFPMA, ABPI, BIA, Bio and DCVMN on Delivering the 100 Days Mission.” GOV.UK, 8 Mar. 2022, https://www.gov.uk/government/publications/joint-statement-on-delivering-the-100-days-mission/joint-statement-from-the-uk-government-cepi-ifpma-abpi-bia-bio-and-dcvmn-on-delivering-the-100-days-mission
Larsen, David A., and Krista R. Wigginton. “Tracking Covid-19 with Wastewater.” Nature Biotechnology, vol. 38, no. 10, 2020, pp. 1151–1153., https://doi.org/10.1038/s41587-020-0690-1
Ma, Cuiping, et al. “A Novel Method to Control Carryover Contamination in Isothermal Nucleic Acid Amplification.” Chemical Communications, vol. 53, no. 77, 2017, pp. 10696–10699., https://doi.org/10.1039/c7cc06469a
Mardis, Connie. Keeping up with POCT Regulatory Compliance - Medical Laboratory Observer. 24 Oct. 2017, https://www.mlo-online.com/information-technology/lis/article/13009284/keeping-up-with-poct-regulatory-compliance
Merindol, Natacha, et al. “Optimization of SARS-COV-2 Detection by RT-QPCR without RNA Extraction.” 2020, https://doi.org/10.1101/2020.04.06.028902.
“Minion.” Oxford Nanopore Technologies, 21 Feb. 2022, nanoporetech.com/products/minion
“MiniSeq System.” MiniSeq Sequencing System | Small, Affordable Benchtop Sequencer, www.illumina.com/systems/sequencing-platforms/miniseq.html
“Miseq System.” MiSeq System | Focused Power for Targeted Gene and Small Genome Sequencing, www.illumina.com/systems/sequencing-platforms/miseq.html
Mustafa, Mujahed I., and Abdelrafie M. Makhawi. “Sherlock and DETECTR: CRISPR-CAS Systems as Potential Rapid Diagnostic Tools for Emerging Infectious Diseases.” Journal of Clinical Microbiology, vol. 59, no. 3, 2021, doi:10.1128/jcm.00745-20
Notomi, T. “Loop-Mediated Isothermal Amplification of DNA.” Nucleic Acids Research, vol. 28, no. 12, 2000, https://doi.org/10.1093/nar/28.12.e63.
“Polymerase Chain Reaction (PCR) Fact Sheet.” Genome.gov, 17 Aug. 2020, https://www.genome.gov/about-genomics/fact-sheets/Polymerase-Chain-Reaction-Fact-Sheet.
“Promethion.” Oxford Nanopore Technologies, 10 Aug. 2022, nanoporetech.com/products/promethion.
Ramuta, Mitchell D., et al. “SARS-COV-2 and Other Respiratory Pathogens Are Detected in Continuous Air Samples from Congregate Settings.” 2022, https://doi.org/10.1101/2022.03.29.22272716.
Shannon, K.E., et al. “Application of Real-Time Quantitative PCR for the Detection of Selected Bacterial Pathogens during Municipal Wastewater Treatment.” Science of The Total Environment, vol. 382, no. 1, 2007, pp. 121–129., https://doi.org/10.1016/j.scitotenv.2007.02.039.
Sheridan, Cormac. “Fast, Portable Tests Come Online to Curb Coronavirus Pandemic.” Nature Biotechnology, vol. 38, no. 5, 2020, pp. 515–518., https://doi.org/10.1038/d41587-020-00010-2.
Soroka, Marianna, et al. “Loop-Mediated Isothermal Amplification (LAMP): The Better Sibling of PCR?” Cells, vol. 10, no. 8, 2021, p. 1931., https://doi.org/10.3390/cells10081931.
“Summary of Probable SARS Cases with Onset of Illness from 1 November 2002 to 31 July 2003.” World Health Organization, World Health Organization, 24 July 2015, www.who.int/publications/m/item/summary-of-probable-sars-cases-with-onset-of-illness-from-1-november-2002-to-31-july-2003.
Tang, Yi, et al. “Advanced Uracil DNA Glycosylase-Supplemented Real-Time Reverse Transcription Loop-Mediated Isothermal Amplification (UDG-RRT-Lamp) Method for Universal and Specific Detection of Tembusu Virus.” Scientific Reports, vol. 6, no. 1, 2016, https://doi.org/10.1038/srep27605
“Truseq RNA Library Prep Kit V2.” TruSeq RNA Library Prep Kit v2 | Simple Libraries from Total RNA, www.illumina.com/products/by-type/sequencing-kits/library-prep-kits/truseq-rna-v2.html.
Venugopal, Anila, et al. “Novel Wastewater Surveillance Strategy for Early Detection of Coronavirus Disease 2019 Hotspots.” Current Opinion in Environmental Science & Health, vol. 17, 2020, pp. 8–13., https://doi.org/10.1016/j.coesh.2020.05.003.
“Voltrax.” Oxford Nanopore Technologies, 23 Mar. 2022, nanoporetech.com/products/voltrax.
Wee, Soon Keong, et al. “Rapid Direct Nucleic Acid Amplification Test without RNA Extraction for SARS-COV-2 Using a Portable PCR Thermocycler.” Genes, vol. 11, no. 6, 2020, p. 664., https://doi.org/10.3390/genes11060664.
Wooley, John C., et al. “A Primer on Metagenomics.” PLoS Computational Biology, vol. 6, no. 2, 2010, doi:10.1371/journal.pcbi.1000667.
Wu, J., Dong, M., Rigatto, C. et al. Lab-on-chip technology for chronic disease diagnosis. npj Digital Med 1, 7 (2018). https://doi.org/10.1038/s41746-017-0014-0
Yu, Lin, et al. “Rapid Detection of Covid-19 Coronavirus Using a Reverse Transcriptional Loop-Mediated Isothermal Amplification (RT-LAMP) Diagnostic Platform.” Clinical Chemistry, vol. 66, no. 7, 2020, pp. 975–977., https://doi.org/10.1093/clinchem/hvaa102.
“2014-2016 Ebola Outbreak in West Africa.” Centers for Disease Control and Prevention, Centers for Disease Control and Prevention, 8 Mar. 2019, www.cdc.gov/vhf/ebola/history/2014-2016-outbreak/index.html.
“2015–16 Zika Virus Epidemic.” Worldwide Outbreak, 7 Mar. 2020, www.worldwideoutbreak.com/blog/cool_timeline/2015-16-zika-virus-epidemic.
Appendices
Appendix 1 – Criteria used for the Weighted Factor Model
Criteria | Explanation |
Current technology | How far are we in having the required technologies to set up such a surveillance system, assuming we can have status quo levels of funding and talent to work on it? |
Cost required | How costly is it to develop such a surveillance system? Amount and level of talent needed would also count as cost in this criteria. |
Scalability | How difficult it is (economically, logistically and politically) to scale such a surveillance system up to a global level? |
Risk | How high are the risks associated with developing this technology, such as the potential of info hazard or technology being misused by bad actors? Risks could also include non-existential forms, such as impact on the ecosystem or causing fragility in the society. |
Timeliness | How quick in theory can irregularities be identified and flagged up using this surveillance system? |
Types of pathogens that can be detected | What is the range of pathogens that can be detected by this surveillance system? Can it detect novel pathogens, or only existing ones? |
Neglectedness | Example questions: Are there many people/groups working on this surveillance system? Does this surveillance system receive much funding? |
Effects across other areas | To what extent do developments in this surveillance method affect other areas that are not related to bio surveillance, or potentially not even related to biosecurity? |
Uniqueness | How easily can the outcomes achieved by this surveillance system be replaced by other forms of surveillance? |
Quality of existing evidence | How much evidence is there now to show that the surveillance system can do what it proposes? |
Appendix 2 - Acknowledgements
First of all, we would like to thank Cambridge Existential Risk Initiative (CERI) for supporting us during this project. In particular, our biosecurity lead Dewi Erwan and our mentor Akhil Bansal.
We are also very grateful for the various experts and organizations that we’ve talked to. These include:


This is a really great summary and reference document, thanks for writing this! I have two comments:
1. There is a difference between detectable levels of RNA/DNA/Antigen and an active, transmissible infection. While most of the time this is not important when thinking about a global surveillance system, it is good to keep in mind especially in PoP testing.
Edit: You address my point 1 in your Part 2
2. You write:
Kevin Esvelt has convincingly argued that we should not do this, because this would publish what and where viruses or pathogens are that could cause a pandemic to bad actors. It seems related to your point on "Potential for Dual-Use by Bad Actors". While the technology itself may not be used by bad actors, the information gathered would immediately be an information hazard which could be used by bad actors.
Again, thanks a lot for this overview of the topic!
Great read, thanks for posting! A quick heads up that many of the links in the table of contents are broken (either linking to start of post, or to non-existent websites).
Summary of this post, and the sequel post Technological Bottlenecks for PCR, LAMP, and Metagenomics Sequencing (feel free to suggest edits!)
Biosurveillance systems help early identification of pathogens that could cause pandemics. The authors weighted existing methods on 10 criteria including usefulness, quality of evidence, feasibility and potential risks.
High scoring methods included: Point-of-person (non-lab tests eg. rapid antigen), clinical (lab tests eg. PCR), digital (reporting cases to a database), and environmental methods (eg. monitoring in wastewater). Technological developments in point-of-person and clinical surveillance (ie. faster, easier, cheaper, home-based tests) is seen as promising. Environmental surveillance would benefit from increasing sensitivity of wastewater testing equipment, and developing new concentration techniques that work for a wide variety of pathogens (bacteria, virus, fungi). Specific bottlenecks and potential solutions (eg. improving performance of LAMP, a cheaper PCR alternative, under cold temperatures) are discussed in the second post.
Slightly lower scoring methods were: animal (frequent sampling and wearable devices) and syndromic (monitoring symptoms). Data sharing between key parties (and preferably cross-country) could assist with syndromic and digital methods. Animal monitoring is less promising as, while 60% of known infectious diseases are zoonotic, we lack the capability to predict virulence and transmissibility to humans.
(If you'd like to see more summaries of top EA and LW forum posts, check out the Weekly Summaries series.)
Interesting read, thank you.
I think your categorisation of surveillance systems could benefit from being done on two axes: the population/environment being tested and the method used for detection. You can basically combine any of these with any of the others, and evaluate their pros/cons orthogonally.
Environments/populations:
Testing technologies:
Thanks for this! I liked the typology of biosurveillance systems. I was confused though by the 'Human / Animal Monitoring' category: why are there humans in this category? If we are testing humans exposed to farm animals, I thought this would fit in the 'sentinel' category. It would make more sense to me if this category was just for testing animals.